Quickstart Example¶
Preamble¶
This is a commented demo file about how to use Cherab. It aims to give an example with the main steps for a basic use of Cherab. We start by importing the classes used in this demo.:
# External imports
import matplotlib.pyplot as plt
import numpy as np
from scipy.constants import electron_mass, atomic_mass
from raysect.core import Vector3D
from raysect.core.scenegraph.utility import print_scenegraph
# Internal imports
from cherab import Species
from cherab.atomic import elements, Line
from cherab.atomic.adas import ADAS
from cherab.distribution import Maxwellian
from cherab.math.mapping.interpolators.interpolators1d import Interpolate1DCubic
from cherab.math.mapping.mappers import IsoMapper3D
from cherab.model.cxs.beaminteraction import CXSBeamPlasmaIntersection
from cherab.model.beam.singleray import SingleRayAttenuator
from cherab_contrib.jet.corentin.ppf_access import *
from cherab_contrib.jet.corentin.data_source import DataSource
from cherab_contrib.jet.corentin.neutralbeam import PINIParameters
from cherab_contrib.jet.corentin.scenegraph import build_jet_scenegraph
In this demo we will simulate the carbon active CX spectra emitted by octant 8 PINIs and seen from different lines of sight. We use input data from pulse 79503 at t=61s.:
PULSE = 79503
TIME = 61.
To achieve this simulation we need to set up the scenegraph, which means placing beams and lines of sight in a first place, and providing parameters and plasma profiles in a second place.
Scenegraph Creation¶
The first step is to generate the scenegraph and to give the geometry and physical models needed. The scenegraph is created through the build_scenegraph function. This function build PINIs and lines of sight with the geometry given in geometry files, which are in cherab.machine.jet.geometries. Given a pulse number, build_scenegraph will look through this module to find the files valid for this pulse and create the scenegraph components whose geometry is defined in these files. For pulse 79503, build_scenegraph will create NIB8 PINIs and ks5 and ks7 lines of sight.
build_scenegraph also need an atomic data provider and the physical models you want to use:
ADAS is the atomic data provider to be used for JET. It is the result of a home-made submodule (cherab.atomic.adas):
atomic_data = ADAS(permit_extrapolation=True)
Physical models: for the beam attenuation model we use SingleRayAttenuator (cherab.model.beam.singleray) and for CX active emissions we use CXSBeamPlasmaIntersection (cherab.model.cxs.beaminteraction) These models must not be called here, as they will be separately for each beam. Instead they must be given as a tuple with the class object in the first place and a dictionary containing arguments in the second place:
attenuation_instruction = (SingleRayAttenuator, {'step': 0.01, 'clamp_to_zero': True})
We want to observe the C5+ lines n=8->7 and n=10->8, so let’s define these lines:
line_1 = Line(elements.carbon, 5, (8, 7))
line_2 = Line(elements.carbon, 5, (10, 8))
The CX active emission model uses this line as a parameter. As it is possible to use different emission models (eg for different lines), they must be given in a list of tuples as defined above:
emission_instructions = [(CXSBeamPlasmaIntersection, {'line': line_1, 'step': 0.03}),
(CXSBeamPlasmaIntersection, {'line': line_2, 'step': 0.03})]
Now we can call build_scenegraph. It returns a World object which is the root of the scenegraph created, a Plasma object which will contain and provide all plasma profiles, and a dictionary named components which contains the main components of the scenegraph with which you will be able to adjust parameters. Here we extract NIB8 (containing the PINIs) and two groups of lines of sight.
world, plasma, components = build_jet_scenegraph(PULSE, atomic_data,
attenuation_instruction,
emission_instructions)
nib8 = components['nib8']
ks5_oct1 = components['ks5_oct1']
ks7_oct8 = components['ks7_oct8']
We can adjust observation parameters for a whole group of lines of sight:
ks5_oct1.min_wavelength = 440
ks5_oct1.max_wavelength = 540
ks5_oct1.spectral_samples = 1000
Plasma Configuration¶
To set up a real plasma, we need to feed plasma with profiles and composition.
Now we are looking for information about the plasma (temperature, density, etc) in the PPF system. Here they are generally given as profile arrays along normalised psi values. As Cherab need any plasma input given as a 3D function the first step is to get normalised psi as a 3D function through the DataSource object:
src = DataSource()
src.time = TIME
src.n_pulse = PULSE
psi = src.get_psi_normalised(exterior_value=1, cached2d=True)
inside = lambda x, y, z: psi(x, y, z) != 1.
psi is the 3D function giving normalised psi (taken from EFIT) inside the LCFS, and an exterior value which can be changed, outside. This outside value can be used to define a inside function returning a boolean, which will be useful to define the plasma later.
To get data from PPF system one can use the python ppf library or a wrapped version of it (cherab.machine.jet.ppf_access) which uses python exceptions instead of error integers (so the content is exactly the same):
ppfsetdevice("JET")
ppfuid('cgiroud', rw='R')
ppfgo(pulse=PULSE, seq=0)
Here a personal DDA is used. We use ppfget to read Data Types as they don’t have a time axis. However one should use ppfgts to read Data Types at a specific time (TIME) if it contains a time axis as Cherab require data at a specific time only:
# normalised psi coordinates
psi_coord = np.array(ppfget(PULSE, 'PRFL', 'C6')[3], dtype=np.float64)
mask = psi_coord <= 1.0 # a mask is created to get only the values inside the LCFS
psi_coord = psi_coord[mask]
flow_velocity_tor_data = np.array(ppfget(PULSE, 'PRFL', 'VT')[2], dtype=np.float64)[mask]
ion_temperature_data = np.array(ppfget(PULSE, 'PRFL', 'TI')[2], dtype=np.float64)[mask]
electron_density_data = np.array(ppfget(PULSE, 'PRFL', 'NE')[2], dtype=np.float64)[mask]
density_c6_data = np.array(ppfget(PULSE, 'PRFL', 'C6')[2], dtype=np.float64)[mask]
# Now these arrays are interpolated to get 1D functions of normalised psi:
flow_velocity_tor_psi = Interpolate1DCubic(psi_coord, flow_velocity_tor_data, extrapolate=True)
ion_temperature_psi = Interpolate1DCubic(psi_coord, ion_temperature_data, extrapolate=True)
electron_density_psi = Interpolate1DCubic(psi_coord, electron_density_data, extrapolate=True)
density_c6_psi = Interpolate1DCubic(psi_coord, density_c6_data, extrapolate=True)
# Extrapolation is allowed by turning to True the `extrapolate` argument.
1D functions are composed with the 3D function psi to get 3D functions giving velocity, temperature, etc. We use IsoMapper3D from a home-made submodule (cherab.math.mapping.mappers) as it is written in cython and then is faster than using lambda functions:
flow_velocity_tor = IsoMapper3D(psi, flow_velocity_tor_psi)
ion_temperature = IsoMapper3D(psi, ion_temperature_psi)
electron_density = IsoMapper3D(psi, electron_density_psi)
density_c6 = IsoMapper3D(psi, density_c6_psi)
So as to be able to handle any velocity profile, The flow velocity must be described as a vector field. Here we just use a toroidal velocity.
flow_velocity = lambda x, y, z: Vector3D(y * flow_velocity_tor(x, y, z),
- x * flow_velocity_tor(x, y, z),
0.) / np.sqrt(x*x + y*y)
In this simulation the plasma is composed of only deuterium and carbon. To get the deuterium density we just have to deduce it from carbon and electron densities:
density_d = electron_density - 6 * density_c6
Note that electron_density and density_c6 can be added and multiplied so easily (they are functions and not values!) because they are returned by IsoMapper3D. Like any function returned by mappers or interpolators submodules they are also written in cython allowing fast evaluation.
Any species (main ion, impurities and electrons) distribution in space and velocity space is described in a Distribution object. Any distribution can be used, but here we only need maxwellian distributions. A Maxwellian function is used to give this distribution directly out of density, temperature, velocity and particle mass (in kg):
d_distribution = Maxwellian(density_d, ion_temperature, flow_velocity,
elements.deuterium.atomic_weight * atomic_mass)
c6_distribution = Maxwellian(density_c6, ion_temperature, flow_velocity,
elements.carbon.atomic_weight * atomic_mass)
e_distribution = Maxwellian(electron_density, ion_temperature, flow_velocity, electron_mass)
Notes:
Here we use the same temperature and velocity for all species, but it is not a requirement at all.
In Cherab, information about species are often given through an Element object from the elements submodule (cherab.atomic.elements). eg the mass of deuterium (in amu) is given by elements.deuterium.atomic_weight.
From here units become important. In Cherab, any density must be given in m^-3, any temperature in eV and any velocity in m/s. These units must be used to create the species distributions.
Now the distributions have been defined, we must associate them to a species. Species are created from an element, a ionisation and a distribution. Electron distribution will be used directly by Cherab so we don’t need to create a species for electrons:
d_species = Species(elements.deuterium, 1, d_distribution)
c6_species = Species(elements.carbon, 6, c6_distribution)
We fill the plasma with the species and the electron distribution we just built, and the inside function for the plasma to know its boundaries.
plasma.inside = inside
plasma.electron_distribution = e_distribution
plasma.set_species([d_species, c6_species])
A plasma is also describe with a magnetic field which must be given (as a 3D vector field, in Tesla). Even if it is not actually used, it is necessary to get ADAS CX rates. Here a toroidal unitary vector field is given:
plasma.b_field = lambda x, y, z: Vector3D(y, -x, 0.).normalise()
PINI Configuration¶
The plasma have been entirely set up, let’s set the PINIs parameters. PINIs will be composed of deuterium:
beam_element = elements.deuterium
PINIs parameters are regrouped under an instance of PINIParameters. For JET octant 8 PINIs, we need information about 8 PINIS, so:
# an array of 8 energies (from PINI1 to PINI8) in eV/amu:
energy = np.array([99707.2, 99707.2, 102295., 102295.,
96386.5, 1., 101416., 101855.], dtype=np.float64) # in eV
energy = energy / beam_element.atomic_weight # in eV/amu
# an array of 3*8 power fractions in W:
powers = np.array([[947863., 1053350., 888242., 1091590., 936611., 0., 1066350., 1066350.], # main component
[550345., 202872., 526211., 213485., 176385., 0., 208979., 208035.], # half component
[324286., 193540., 283783., 190731., 182980., 0., 190921., 187949.]], # third component
dtype=np.float64)
# an array of 8 booleans to give the status (turned on/turned off) of the PINIs:
turned_on = np.array([True, True, True, True, False, False, True, False], dtype=bool)
# Note PINI6 has 0 power and an unusual energy, but these values will not be
# used as PINI6 is turned off.
# Regroupment of data, including the beam species:
pini_parameters = PINIParameters(energy, powers, turned_on, beam_element)
# We can feed NIB8 with these PINI parameters:
nib8.pini_parameters = pini_parameters
Observation¶
The scenegraph have been completely set up now! To be sure, we can have a look at it with print_scenegraph from raysect.core.scenegraph.utility:
print_scenegraph(world)
To make observations, we must choose a line of sight or a group of lines of sight and call the observe method. The display method allow to plot the result.
From a line of sight group, each line of sight can be accessed by calling get_los with the name of the line of sight. Names are defined in the geometry files.
los_o8l10 = ks7_oct8.get_los('O8L_10')
ks5_oct1 is a particular case as it is a group of groups. Each of its group has a name which is defined in its geometry file too (they represents C, D, B from 1 to 3 and B from 4 to 6 lines of sight). You can access a particular group with get_los_group method:
los_d5 = ks5_oct1.get_los_group('D Lines').get_los('D5')
# Or get directly a particular line of sight:
los_d5_direct = ks5_oct1.get_los('D5')
print(los_d5 is los_d5_direct)
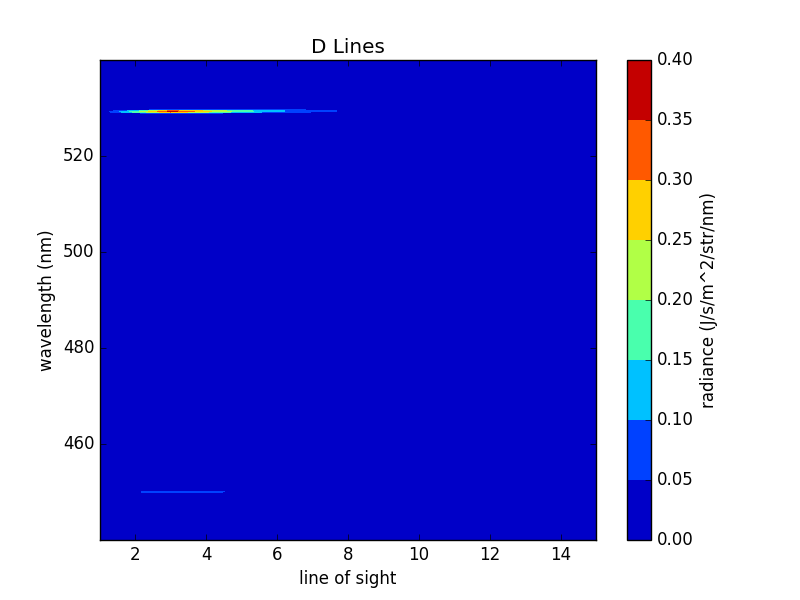
For instance let’s make all the D lines from ks5_oct1 observe and display the result:
ks5_oct1.get_los_group('D Lines').observe()
ks5_oct1.get_los_group('D Lines').display()
If you want to have a look at a particular spectrum, eg the one of D5:
los_d5.display()
# the display is by default in J/s/m^3/str/nm, it can be turned in photons/s/m^3/str/nm:
los_d5.display(unit='ph')
After observing, any line of sight store the measured spectrum in a spectrum attribute:
d5_spectrum = los_d5.spectrum
plt.plot(d5_spectrum.wavelengths, d5_spectrum.samples)
plt.show()
As for the display, the spectrum is in J/s/m^3/str/nm, you can generate an array of data in photons/s/m^3/str/nm using to_photons method:
photons_samples = d5_spectrum.to_photons()
plt.plot(d5_spectrum.wavelengths, photons_samples)
plt.show()